 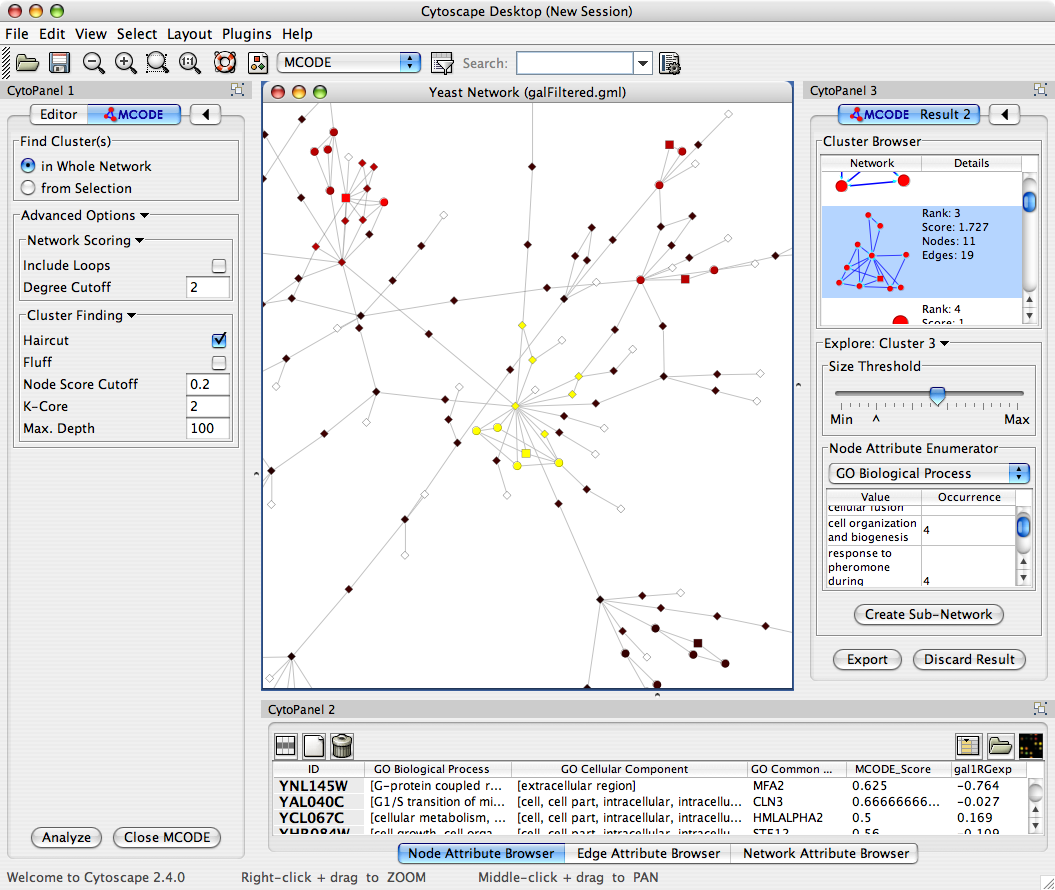
of nativ ways to save graphs, for example Cytoscape, which well see later. Protein contact network (PCN) formalism allows for a rational, structure based approach to both the drug action modes. First possibility to represent graphs is through their adjacency matrix. In the case of an allosteric (non-local) paradigm, the focus shifts toward the signal transmission across the protein molecule: this renders the 'most promising' binding sites those residues with the most 'central' position that have the higher probability, when perturbed by a ligand, to generalize the perturbation to the entire structure. Given the interaction with other mac-romolecules is the core of protein physiological role, in a local (orthosteric) paradigm of pharmacological action, we maximize the probability of perturbing the system by an agent binding in the same place where such interaction takes place: the most flexible parts of the structure. Cytoscape app that enables AdjacencyMatri圎xport formats for directed & undirected networks. Conclusion: This stems from the large amount of observations pointing to natively unfolded tracts of protein sequences as responsible for protein-protein interactions. 
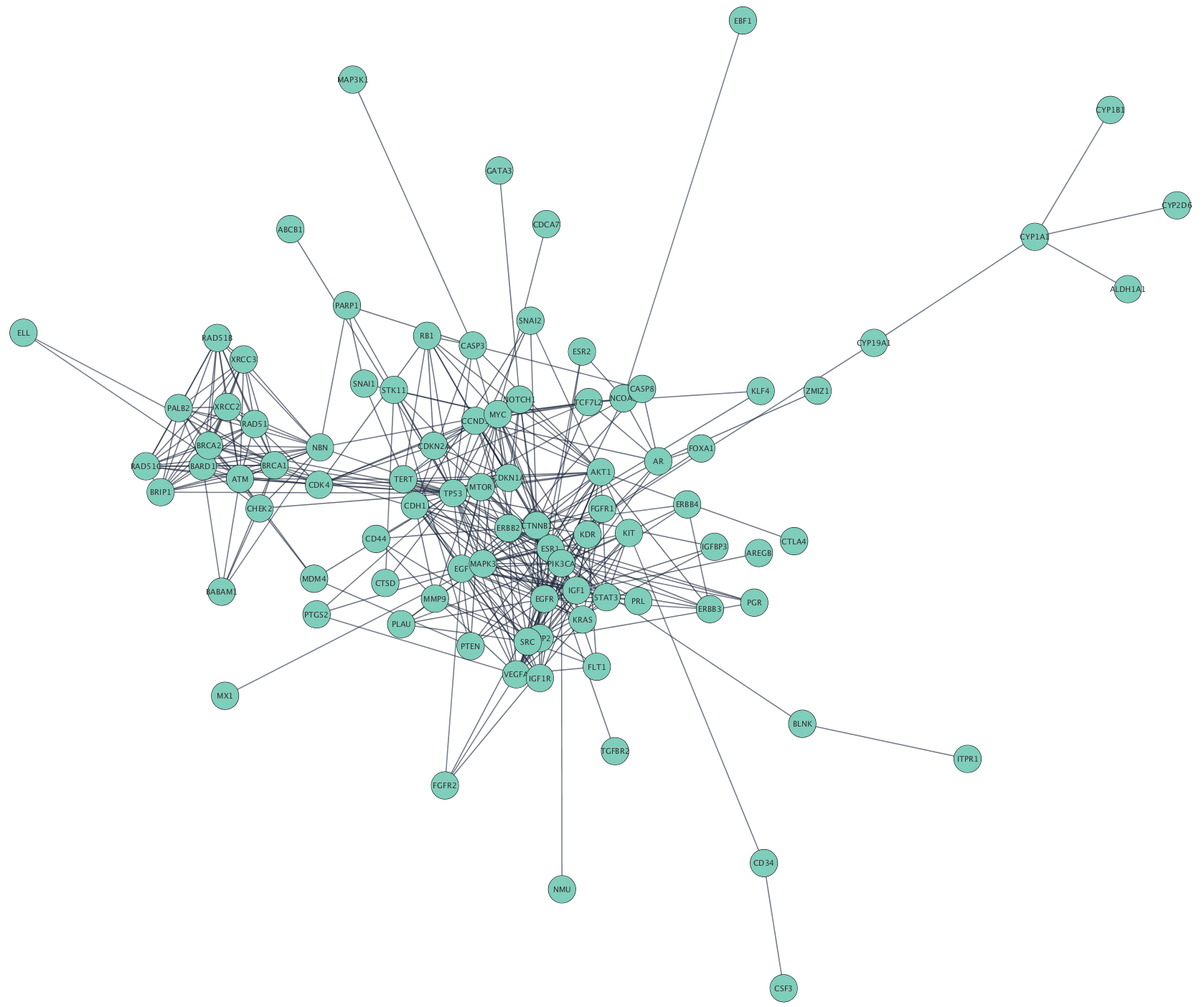
In the orthosteric paradigm, the 'hot spots' for ligand binding are those residues endowed with higher flexibility. In this scenario, the link between relative flexibility and druggability of protein targets has two very distinctive faces depending upon the orthosteric/allosteric paradigm of drug action. Much is known about the binding of small molecules to single pep-tide chains, but much more is still to be discovered about macromolecular complexes and how binding can be affected by modulators. Hence, targeting protein-protein interfaces is actually the " holy grail " of pharmacology. to Cytoscape-Discuss Hi, There is no export as an adjacency matrix available, but calculating the network paramters can be done using the NetworkAnalyzer tool as described here. The network was illustrated in the form of nodes and edges, where each node represents a single gene, and each edge. Background: Protein-protein interactions embody the main target of drug discovery due to their vast presence in physiological mechanisms. The adjacency matrix was exported into Cytoscape via the aMatReader plugin 28.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
